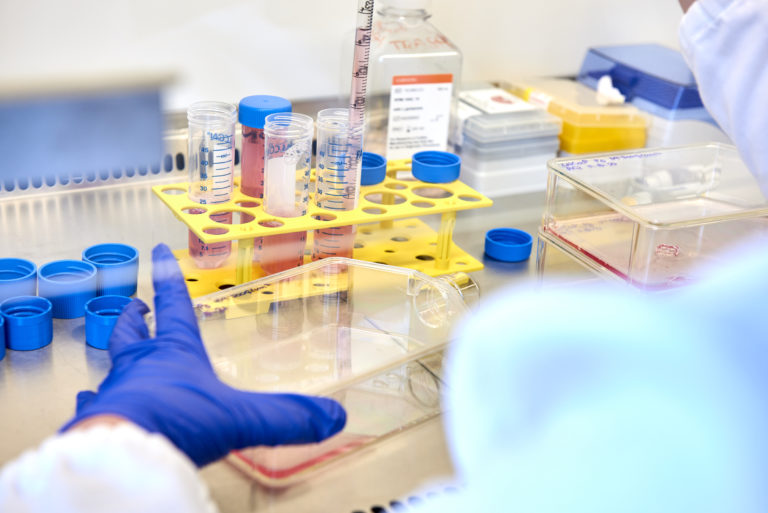
Nematic Protein Organization Technique (NPOT®) is a groundbreaking, label-free platform that isolates and identifies the drug-induced interactome; the dynamic network of proteins interacting directly and indirectly with a therapeutic compound. NPOT® operates directly in native biological matrices, including cell lines, human/animal tissue homogenates, and pathological samples (e.g., biopsies), preserving physiological protein conformations and interactions. Compatible with small molecules, therapeutic antibodies (mono-/multi-specific), peptides, and high molecular weight compounds, NPOT® delivers unparalleled insights into drug behaviour for precision medicine and therapeutic development.
Key deliverables of NPOT® analysis
Primary & secondary targets
Identification of on-target, off-target, and polypharmacological interaction.
Drug interactome
Mapping of effector proteins, co-regulators, and signaling complexes.
Mechanism of action (MoA)
Elucidation of drug-induced signaling pathways and functional networks.
Biomarkers
Discovery of biomarkers linked to drug response, resistance, or patient stratification.
Key advantages of NPOT®
Physiological relevance
Maintains native quaternary structures and protein-protein interactions by replicating physiological/pathological environments, enabling target validation in native conformations (e.g., GPCR dimers, membrane-embedded receptors).
Comprehensive analysis
Bridges the gap between in vitro assays and in vivo complexity by studying drug-protein and protein-protein interactions directly in pathological tissues (e.g., biopsies), ensuring observations reflect true in vivo conditions. This allows researchers to:
- Decipher polypharmacology and off-target effects in disease-relevant contexts.
- Accelerate translational research using clinical samples.
Specificity
Employs parallel negative controls (e.g., inactive enantiomers) to distinguish specific interactors from non-specific background proteins, ensuring robust, artifact-free results.
How does NPOT® work?
NPOT® is founded on the principles of Kirkwood-Buff molecular crowding and aggregation theory, enabling the formation and label-free identification of physiologically and pathologically relevant macromolecular complexes. The process unfolds in the following detailed steps:
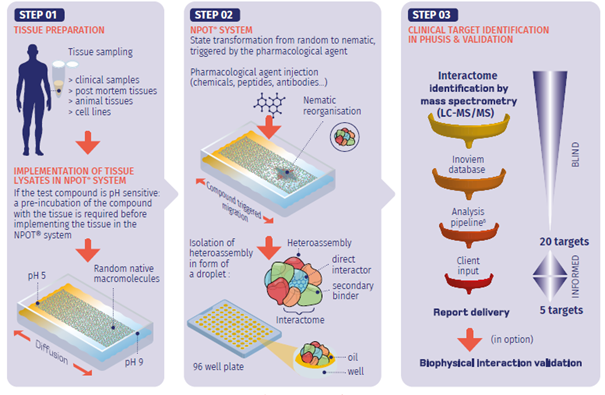
Biological matrix sample processing
Cells and tissue homogenates are subjected to mechanical disruption (e.g., freeze-thaw cycles) in the absence of detergents or reducing agents. This careful processing helps to preserve native conformations and protein-protein interactions—crucial for the functionality of receptors (such as GPCR dimerization) and other macromolecular assemblies.
Implementation of a physiological pH gradient
The homogenates are introduced into a continuous, transitory, and reversible pH gradient (pH 5–9), established using ampholytes and formulated at an osmolality matching the extracellular environment. This gradient replicates the physiological diversity of pH found in cellular compartments (ranging from acidic lysosomes to basic mitochondria), ensuring that proteins and protein complexes migrate naturally to their pH-specific equilibrium based on their charge, molecular volume, and energy (ΔG). The pH gradient is critical for the proper formation and isolation of the drug interactome.
For an in-depth explanation of the pH gradient’s role, please visit this page.
Test compound introduction
Upon addition, the test compound interacts with its target proteins, altering their surface charge distribution. This interaction shifts the protein complexes to their zwitterionic point (net charge = 0), resulting in increased hydrophobicity and the specific precipitation of both direct and indirect interactors. The formation of a focal nematic viscosity within the pH gradient is indicative of interactome assembly, typically completed within 3–9 hours. Specificity is ensured via parallel negative controls (inactive enantiomers/analogs), eliminating nonspecific background.
Isolation of the interactome
The precipitated interactome is extracted into a mineral oil-phase solution, preventing resolubilization. This precipitation is strictly dependent on the presence of the active compound—no precipitation is observed in the absence of the drug—ensuring a high degree of specificity in the interactome isolation.
Gel stacking and filtration
The isolated proteins are subsequently solubilized and subjected to SDS-based gel stacking filtration. This step is followed by enzymatic digestion (e.g., trypsin) to generate peptides suitable for downstream proteomics sequencing.
LC-MS/MS sequencing
The peptides derived from the interactome are sequenced using state-of-the-art proteomic platforms (e.g., Bruker TIMS TOF pro 2), ensuring high resolution and sensitivity.
Why NPOT® for translational drug development?
At Inoviem, our NPOT® platform stands at the nexus of cutting-edge technology and clinical relevance, providing an exhaustive toolset for drug development services.
Native context
No detergents, no overexpression – study targets in pathological tissue.
Functional specificity
Negative controls and InOPERA® eliminate false positives.
Translational power
Direct analysis of clinical samples accelerates biomarker discovery.