- Services
Services
Inoviem provides a full range of services from drug discovery to clinical development by leveraging its platforms and direct access to human pathological specimens.
All the platforms and services are label-free with no modification of the original molecules.Unravel disease relevant Mode of Action (MoA) of your molecule directly on human tissues.
Identify and validate targets in your study model or in Human.
Thanks to our ex-vivo pharmacology expertise we identify the best disease and patients subgroups.
Using patients’ samples and PIMS® technology, we identify those individuals who can benefit best from your compound as soon as the early stage of preclinical studies.
Compounds of interest can be profiled for their drugability based on their mode of action and efficacy.
Translational pharmacology empowers your ability to identify biomarkers and advance clinical developments.
Advanced SPR and proteomics drive precise analysis of biomolecular interactions and protein landscapes.
Meticulous collection and biobanking of human samples fuel robust, translational research.
- Platforms
Platforms
Inoviem provides breakthrough protein technologies for each phase of the drug development process.
One of the major advantages of our label-free technologies used under physiological conditions and in human tissue – NPOT® and PIMS® – is that the test compound retains its original structure, as well as the exact same molecular structure that would be used in therapyPIMS®
Designed to study molecular interactions and predict therapeutic responses in biomedical research
Biomarkers discovery
Methodology for the proteomic profiling of human individuals in the context of translational evaluation of biomarkers and clinical development of new drug candidates.
- Knowledge centre
- About us
About us
Inoviem Scientific is a Bioanalytical R&D company established in 2011 in Strasburg – France.

Our mission
Our mission is clear: to get the right treatment to the right patients within the right therapeutic window.
- Contact us
Proteomic profiling for biomarkers discovery in human whole blood and complex human matrixes (serum, plasma, sputum, balf, urine)
Inoviem has developed a sensitive and accurate proteomic methodology for the proteomic profiling of human individuals in the context of translational evaluation of biomarkers and clinical development of new drug candidates. The methodology works with both human frozen whole blood and any other matrixes including plasma/sera, urine, sputum etc.
The workflow of MS-based proteomics experiments includes the following steps: experimental design, sampling, tissue/cell preparation, protein extraction, separation, MS analysis, protein identification, statistical analysis of data, validation of identification, quantification, and data analysis. The experimental design is detailed in a table, which is submitted for approval.

Method for frozen whole blood: 3D fractioning sample preparation
This methodology has the great potential of analyzing frozen whole blood for biomarkers discovery by extracting hemoglobin. Haemoglobin indeed poses a significant challenge in whole blood proteomic analysis due to its high abundance and potential to interfere with the detection of other proteins. Inoviem developed a label free, non-chromatography-based methodology to extract the haemoglobin and to elute the proteins bound to albumin.
The steps for whole blood preparation:
Haemoglobin depletion
Iron ions (Fe2+) are replaced by Mn2+ and Mg2+ with gradually increasing saline forces. This will precipitate free hemoglobin and induce hemagglutination.
Charge distribution-based fractioning
Stepwise change in ionic forces from neutral to acid between with pH interval 4 to 6.
Hydrophobicity-based fractioning
Separation as well as isolation of hydrophobic and hydrophilic proteins using proprietary three phases gradient. Each phase is analyzed thereafter in triplicate.
Size-based fractioning
Filtration on SDS-PAGE gradient gel, isolation of proteins and proceed to proteomics.
Each fractioned phase is sequenced via label-free quantitative proteomics (LC-MS/MS) and the proteins are analyzed both quantitatively and qualitatively.
This method identifies an average of 710 proteins per blood sample, outperforming other depletion and fractionation methods available on the market, which typically rely on labelling and conjugation chemistry approaches.
Qualitative analysis
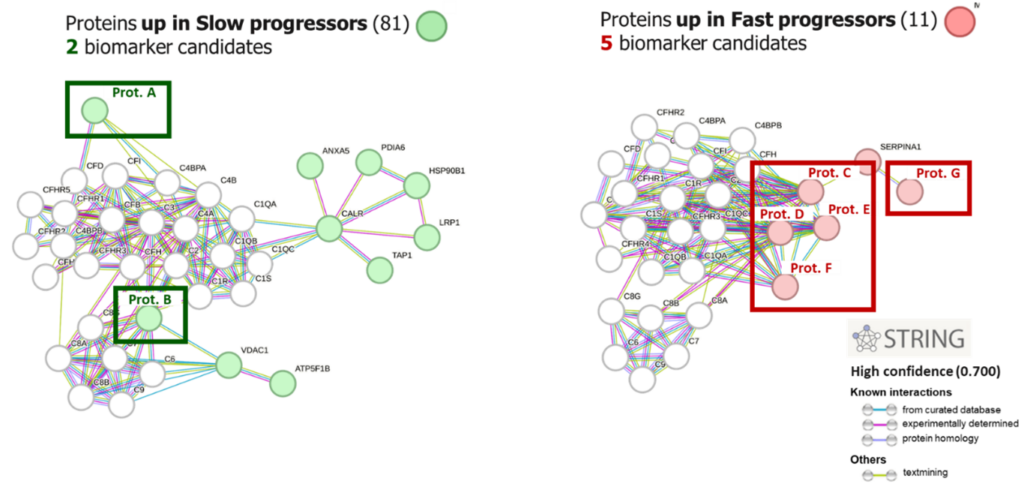
Label free proteomics analysis reveals many proteins that can be easily sorted and ranked by Inoviem’s InOpera database, able to discriminate between frequent and specific proteins within the dataset. This opens the possibility to perform pathway enrichment analysis aiming to the identification of modulated pathways related to disease features (e.g. progression rate, stage) or to treatment response (e.g. response/non-response related pathways) leading to the identification of potential biomarkers candidates.
Quantitative analysis
Each fractioned blood sample undergoes to label-free quantification in order to get an exhaustive proteomic signature of each patient.
Analyses are based on two steps:
- Global differences analysis
- Protein distribution (cell location)
- Protein’s physico-chemical properties (hydrophobicity, charge distribution, aliphatic and Boman indexes)
- Identified peptides (numbers/protein for each group)
- Specific protein analysis
- Enriched/impoverished proteins in patient groups
- Protein/protein interaction and network structure deciphering
- Relevance with clinical data
These types of analyses can reveal modulated proteins based on patient’s group therefore revealing potential biomarkers candidates. With this methodology we can get the following types of biomarkers:
- Disease progression biomarkers: Useful for monitoring the course of the disease.
- Prognostic biomarkers: Useful for predicting the likely outcome or course of the disease.
- Diagnostic biomarkers: Useful for identifying the presence of a disease.
- Therapeutic biomarkers: Useful for predicting response to treatment and personalizing therapy.