- Services
Services
Inoviem provides a full range of services from drug discovery to clinical development by leveraging its platforms and direct access to human pathological specimens.
All the platforms and services are label-free with no modification of the original molecules.Unravel disease relevant Mode of Action (MoA) of your molecule directly on human tissues.
Identify and validate targets in your study model or in Human.
Thanks to our ex-vivo pharmacology expertise we identify the best disease and patients subgroups.
Using patients’ samples and PIMS® technology, we identify those individuals who can benefit best from your compound as soon as the early stage of preclinical studies.
Compounds of interest can be profiled for their drugability based on their mode of action and efficacy.
Translational pharmacology empowers your ability to identify biomarkers and advance clinical developments.
Advanced SPR and proteomics drive precise analysis of biomolecular interactions and protein landscapes.
Meticulous collection and biobanking of human samples fuel robust, translational research.
- Platforms
Platforms
Inoviem provides breakthrough protein technologies for each phase of the drug development process.
One of the major advantages of our label-free technologies used under physiological conditions and in human tissue – NPOT® and PIMS® – is that the test compound retains its original structure, as well as the exact same molecular structure that would be used in therapyPIMS®
Designed to study molecular interactions and predict therapeutic responses in biomedical research
Biomarkers discovery
Methodology for the proteomic profiling of human individuals in the context of translational evaluation of biomarkers and clinical development of new drug candidates.
- Knowledge centre
- About us
About us
Inoviem Scientific is a Bioanalytical R&D company established in 2011 in Strasburg – France.

Our mission
Our mission is clear: to get the right treatment to the right patients within the right therapeutic window.
- Contact us
NPOT®
Unlocking drug mechanism of action in native biological contexts
NPOT® (Nematic Protein Organization Technique) is a biophysical platform that maps how drugs interact with proteins in their native, unaltered state. By analysing tissues, cell lines, or patient biopsies directly, NPOT® identifies drug targets, interaction networks, and biomarkers – without disrupting the natural biological environment. Compatible with small molecules, antibodies, peptides, and large compounds, NPOT® accelerates drug discovery by revealing how therapies truly work in disease contexts.
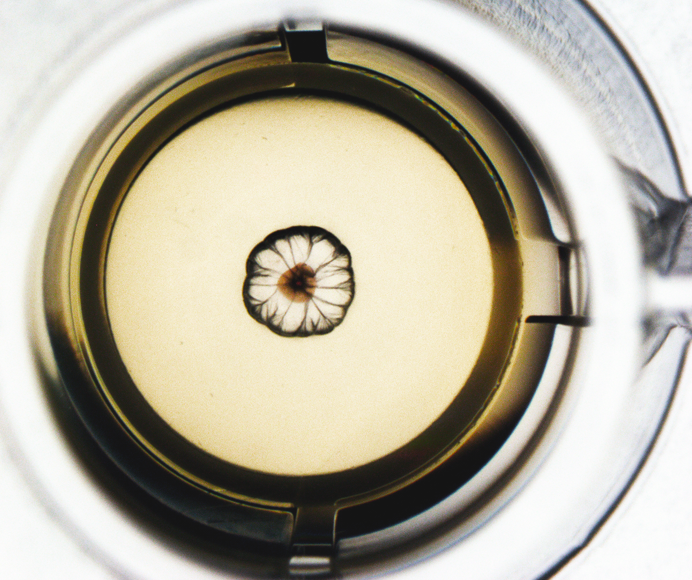

The concept
Leveraging Kirkwood-Buff molecular crowding and aggregation theory, NPOT® harnesses the power of any chemical entity—be it a small molecule, protein, or high molecular weight compound—to shift the equilibrium of cellular proteins in a pH gradient, reflecting different cell compartments. This shift triggers the formation of a heteroassembly, which is then isolated and analyzed via LC-MS/MS.
Next, we use InOPERA®, Inoviem Scientific proprietary database, to distinguish nonspecific proteins (which display a high frequency of appearance within database integrated biosamples), from specific proteins, which include the target candidate.
NPOT® is a game-changer, offering an unbiased approach that not only identifies targets but also reveals the early stages of the mode of action (MOA). Proven effective across nearly all therapeutic areas, NPOT® pinpoints direct drug targets (ON- and OFF-targets) and indirect effectors, uncovering pathways and MOAs. Each identified protein undergoes thorough multidirectional analysis, including functional studies, physico-chemical property assessments, and UniProt annotation.
To ensure precision, NPOT® includes a parallel analysis with a negative control compound, such as an inactive enantiomer or a structurally similar compound with minimal pharmacological activity. This critical step guarantees the identification of selective and specific proteins within the interactome, setting a new standard for drug discovery and development.
Key deliverables:
- Drug Targets: On-target, off-target, and polypharmacological interactions.
- Interactome Map: Proteins and pathways directly or indirectly engaged by your dr
- Mechanism of Action: How your compound behaves in complex biological systems.
- Biomarkers: Clinically relevant markers for patient stratification or treatment response.

How NPOT® Works
NPOT® combines biophysical principles with advanced proteomics to study drug effects in native tissues:
- Sample preparation: Tissues/cells are gently processed to preserve natural protein interactions.
- Native Environment Setup: Proteins migrate in a pH environment mirroring human cells.
- Compound interaction: The drug triggers protein aggregation, isolating its unique interactome.
- Analysis: Precipitated proteins are identified via cutting-edge proteomics and validated using Inoviem’s proprietary tools.
Key deliverables:
Relevance
Works in human tissues, including biopsies—no artificial models.
Specificity
Uses inactive compounds as controls to filter out false positives.
Speed
Go from sample to actionable insights in days, accelerating translational research.
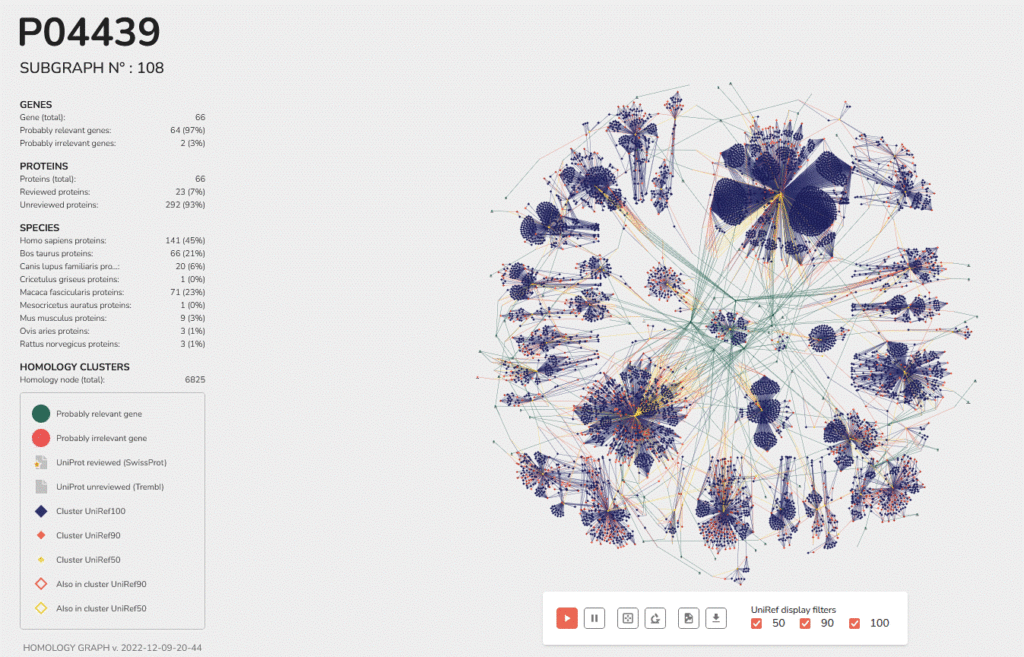
Data Analysis and Interpretation with InOPERA®
Following LC-MS/MS sequencing, Inoviem’s proprietary software (InOPERA®) rigorously mines the data to reveal true interactome members. Our InOPERA® database cross-references each protein against more than 300 NPOT® studies, using frequency analysis to exclude nonspecific ontaminants (proteins that appear in over 25% of studies, such as actin).
In parallel, we conduct detailed physicochemical profiling – comparing
hydrophobicity (GRAVY index), net charge, and interaction potential (Boman index) between test compounds and controls – to prioritize selective interactors. Combined with functional annotations from resources like UniProt, this comprehensive workflow provides an unbiased, complementary approach for pinpointing the primary targets of drug candidates.
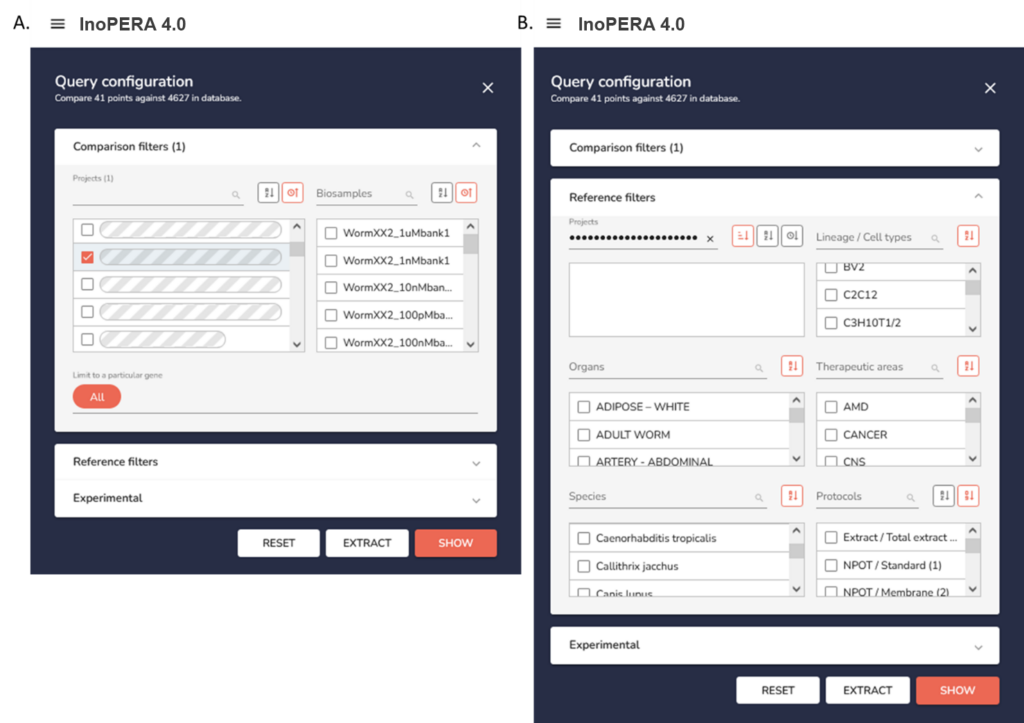
InOPERA® application user interface. A Comparison filter. Selection of imported dataset(s) and/or selection of biosample(s) to be compared in the InoPERA® database. B. Selection of the reference filter(s) to define the subset of the database used for frequency calculation. Different reference filter strategies can be combined, based on the cell types, organs, therapeutic areas, species, and protocols.

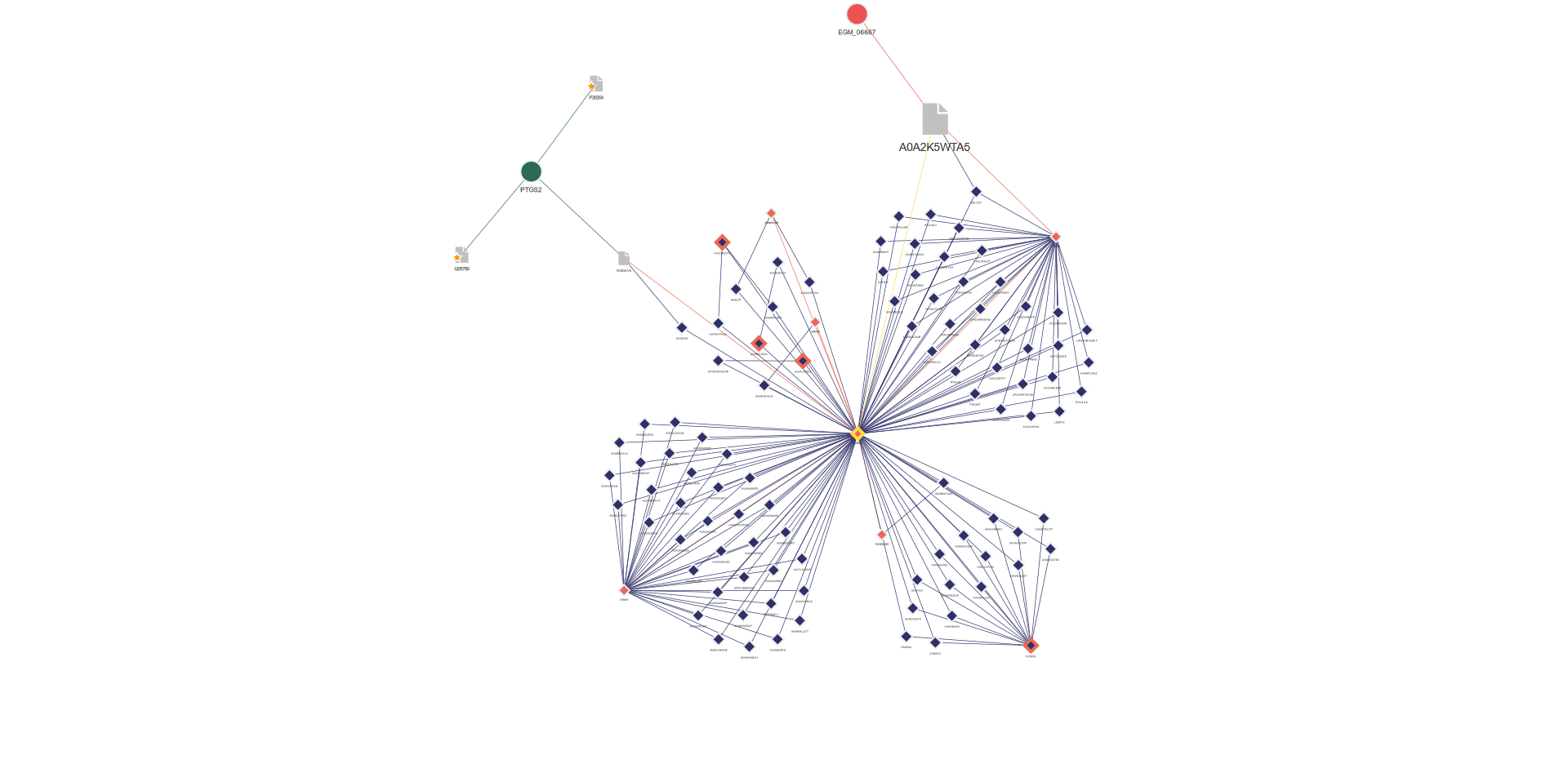
Pathway enrichment and functional analysis
The proteins enriched by NPOT® are further interrogated using multiple functional enrichment analysis tools. These tools integrate data across various species and validation methodologies, associating the identified proteins with biological functions and pathways. Complementary network analyses further enhance our understanding by visualizing the functional networks and elucidating the interplay among interactome components:
- STRING database integration: Protein networks are mapped to biological pathways (e.g., kinase signaling, ubiquitination) and visualized in functional clusters.
- Multi-omics annotation: Enriched proteins are linked to disease pathways, gene ontology (GO), and clinical biomarkers via UniProt, KEGG, and Reactome.
Pathway and target validation
Post-analysis, Inoviem validates both target candidates and the identified protein-protein interactions using Surface Plasmon Resonance (SPR). This validation step is applicable to both membrane-bound and cytosolic proteins, allowing for precise measurements of binding specificity, active concentration, and the kinetic and thermodynamic parameters of the interactions.
By preserving the native state of biological systems, NPOT® ensures that insights into drug action and target engagement are as close to the in vivo reality as possible, thus driving precision medicine forward.
For scientists, by scientists
NPOT® bridges the gap between lab experiments and real-world biology. Whether you’re optimizing lead compounds or de-risking clinical candidates, NPOT® delivers mechanistic clarity for smarter decisions.
